CHAPTER 4: EXTENSIONS OF MENDELIAN INHERITANCE.
Many crosses do not yield simple Mendelian ratios. Instead, the ratios are modified.
These modifications reflect complexities in gene expression not complexities in
inheritance of the genes. In other words, the genes (and genotypes) are still inherited
according to Mendelian rules, but the phenotypes do not conform to simple Mendelian rules.
I. Variations of Dominance
A. Incomplete dominance
1. Phenotype of the heterozygote is intermediate between the phenotypes of the
homozygotes
snapdragon plant with red petals (C1C1) x plant with white petals
(C2C2)
1/4 red C1C1
1/2 pink C1C1
1/4 white C2C2
B. Co-dominance
1. Phenotype of the heterozygote shows the phenotype of both homozygotes.
For example, the gene required for production of a glycoprotein on the surface of red
blood cells (basis for the M and N blood groups).
LM LM has only M antigens on the surface
LN LN has only N antigens on the surface
LM LN has both antigens on the surface.
C. The distinction between incomplete dominance and codominance is dependent upon the
phenotypic "level" to which you are referring (i.e. sickle cell anemia example
pg. 92-93 of your textbook).
II. Multiple Alleles
A. Although an individual organism (say you) can have only two alleles of a gene
(because there are only two chromosomes), among a population (say the entire class), there
may be multiple alleles which are said to be in an allelic series
B. For example, the genes that are responsible for the synthesis of sugars bound to
a lipid on the surface of the red blood cells (basis for another type of human blood
groups)
3 alleles (i or IA or IB) but any individual can only have two of
them.
| Blood Group |
Genotype |
Phenotype of sugar on surface |
| O |
ii |
no sugar on surface |
| A |
IAIA or IAi |
N-acetyl galactosamine |
| B |
IBIB or IBi |
galactose |
| AB |
IAIB |
N-acetyl galactosamine and galactose |
IA and IB are dominant to i but codominant to each other.
C. Other allelic series
C - = full color - dark gray
cch = chinchilla - pale gray
ch = himalayan - albino with dark extremities
c = albino
Dominance = C >cch>ch>c
D. Test for Allelism
How does one know that a set of contrasting phenotypes is determined by alleles of only
one gene (vs. 2 sets of genes that influence each other which is described in Section IV)
Test for allelism and simple dominance: Observation of F2 ratios (3:1) from crosses of
all pairwise combinations of pure breeding strains.
III. Lethal Alleles
A. For example,
yellow mice x normal pure breeding agouti type mouse ---à
1/2 yellow and 1/2 normal
Therefore, yellow was dominant and parent mouse was heterozygous (because agouti was
pure breeding)
yellow mice x yellow mice ---à 2/3 yellow mice and 1/3
agouti
A homozygous dominant mouse (pure breeding) could never be obtained.
Since ratio expected from heterozygous cross is 1:2:1, this suggested that homozygous
dominant was being eliminated -- lethal – to alter the ratio to 2:1.
The allele causing yellow color is dominant for coat color. However, in terms of
viability, the allele is recessive. This the alleles is pleotrophic (affects two or more
distinct traits).
Use these symbols since it is dominant for color mutant but recessive for lethality.
AY A x AY A
1/4 AYAY lethal
2/4 AY A yellow
1/4 A A normal color
B. Can have recessive lethal alleles (as shown above) and also recessive
dominant lethal alleles (where heterozygote is lethal too)
C. Lethal alleles illustrate the idea of the pleotrophic gene (gene that results
in more than one phenotypic effect)
D. Can also have semilethal genes where not every individual with the lethal
genotype dies.
IV. Several Genes Affecting the Same Character
The cellular functions of products encoded by more than one gene affect the
phenotype of one trait. The key to determining that you have one trait controlled by more
than one gene is modified (i.e. not 3:1) Mendelian ratios in a cross of 2 heterozygous
individuals. Ratios are typically in 16ths if two genes are involved.
A. Simple dominance with no epistasis and with each loci independently
influencing the trait: Phenotypic ratio is 9:3:3:1
For example, coat color in mice.
Gene A determines the distribution of pigment in the hair in mice:
A- = agouti (yellow band on each hair)
aa = no yellow agouti band, solid color
Gene B determines the color of the pigment:
B- = black and bb = brown
Cross AABB (wildtype agouti) x aabb (brown) ---------à
F1 are all AaBb (wildtype agouti)
Self the F1s (AaBb) ------à
9/16 A-B- agouti
3/16 aaB- cinnamon
3/16 A-bb black
1/16 aabb brown
In this case we have 4 different phenotypes represented for the trait of hair color.
B. Epistasis: A situation where the alleles of one gene mask the phenotype of
another gene. For epistatic cases, one observes a reduction in the number of phenotypic
classes expected.
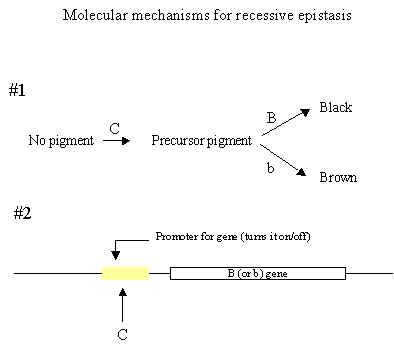
1. Recessive Epistasis: Phenotypic ratio is 9:3:4
a) For example, mouse coat color again
C gene allows the formation of coat pigments.
C- = pigment formation cc = no pigment formation (albino)
Gene B determines the color of the pigment:
B- = black and bb = brown
Cross BBCC (black) x bbcc (albino) ---------à
F1 are all BbCc (black)
Self the F1s (BbCc) ------à
9/16 B-C- black
3/16 bbC- brown
3/16 B-cc albino
1/16 bbcc albino
So overall phenotypic ratios were
9/16 B-C- black
3/16 bbC- brown
4/16 --cc albino
b) Any genotype that has cc (ie either B-cc or bbcc) will be albino; the homozygous
recessive cc alleles are epistatic to the alleles at the locus. This is recessive
epistasis.
c)
2. Dominant epistasis: Phenotypic ratio is 12:3:1
For example, fruit color in squash
Gene a allows the formation of color.
A- = no color formation aa = color formation
Gene B determines the color
B- = yellow and bb = green
Cross AABB (white) x aabb (green) ---------à
F1 are AaBb (white)
Self the F1s (AaBb) -------- -à
9/16 A-B- white
3/16 A-bb white
3/16 aaB- yellow
1/16 aabb green
Thus the total phenotypic ration is
12/16 white
3/16 yellow
1/16 green
3. Duplicate recessive epistasis (AKA complementary gene action): Phenotypic
ratio is 9:7
a) For example, flower color in sweet peas:
Gene A is required for the formation of color.
A- = purple aa = white
Gene B is also required for the formation of color.
B- = purple bb = white
Crossed 2 pure breeding plants that produced white flowers
AAbb X aaBB ---------------à
F1 were all unexpectedly purple (AaBb)
Self the F1s (AaBb) -------- -à
9/16 A-B- purple
3/16 A-bb white
3/16 aaB- white
1/16 aabb white
Thus the total phenotypic ratio is
9/16 purple
7/16 white
b) Explanation was that white flowers in each line were a result of homozygous
recessive alleles in two separate genes. Thus the F1 were all purple because the genotype
was AaBb
Precursor (colorless) –Aà
Intermediate (colorless) –Bà Purple
c) Complementation can be used to determine whether different variants with the
same phenotype contain mutations in the same gene.
C. Suppression: One gene cancels the expression of an abnormal allele thereby
RETURNING IT TO WILD TYPE PHENOTYPE.
Please note that this is different that epistasis. The epistatic allele introduces a
third phenotype (except for complementary gene action), whereas the suppressor allele
simply results in a wildtype phenotype even though the genotype suggests that the
phenotype should be mutant.
1. Recessive suppression of a recessive allele: Phenotypic ratio is
13:3
For a hypothetical gene B
where B- = wt and bb = mutant,
consider a recessive suppressor where
A- = no suppression and aa = suppression of bb
Selfing heterozygotes (AaBb) -------- -à
9/16 A-B- wt
3/16 A-bb mutant
3/16 aaB- wt
1/16 aabb wt (due to suppression)
Thus the total phenotypic ration is
13/16 wt
3/16 mutant
In Drosophila, the recessive allele pd results in purple eye color
Purple eye color can be suppressed by the recessive allele su
pdpd SuSu X PdPd susu --à F1 were all Pdpd Susu
Selfed the F1 -à
9/16 Pd- Su red
3/16 pdpd Su- purple
3/16 Pd- susu red
1/16 pdpd susu red (due to suppression)
Thus the overall phenotypic ratios are
13/16 red (wt)
3/16 purple (mutant)
2. Dominant suppression of a dominant allele: Phenotypic ratio is 13:3
For a hypothetical gene B
where B- = mutant and bb = wt,
consider a dominant suppressor where
A- = suppression of B- and aa = no suppression
Selfing heterozygotes (AaBb) -------- -à
9/16 A-B- wt (due to suppression)
3/16 A-bb wt
3/16 aaB- mutant
1/16 aabb wt
Thus the total phenotypic ration is
13/16 wt
3/16 mutant
3. Recessive suppression of a dominant allele: Phenotypic ratio is 9
mutant: 7 wt
For a hypothetical gene B
where B- = mutant and bb = wt,
consider a recessive suppressor where
A- = no suppression and aa = suppression of B-
Selfing heterozygotes (AaBb) -------- -à
9/16 A-B- mutant
3/16 A-bb wt
3/16 aaB- wt (due to suppression)
1/16 aabb wt
Thus the total phenotypic ration is
9/16 mutant
7/16 wt
4. Dominant suppression of a recessive allele (this is just another way
of saying duplicate gene action): Phenotypic ratio is 15:1
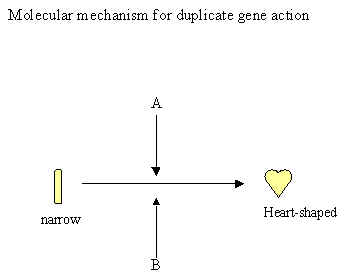
D. Duplicate gene action: Phenotypic ratio is 15:1
1. For example, fruit shape in the plant shepherd’s purse
Gene A determines the shape of the fruit
A- = heart shaped fruit aa = narrow fruit
Gene B also can determine the shape of the fruit
B- = heart shaped fruit bb = narrow fruit
Crossed 2 pure breeding plants that produced white flowers
AABB (heart shaped) X aabb (narrow) ---------------à
F1 were all heart shaped (AaBb)
Self the F1s (AaBb) -------- -à
9/16 A-B- heart shaped
3/16 A-bb heart shaped
3/16 aaB- heart shaped
1/16 aabb narrow
Thus the total phenotypic ration is
15/16 heart shaped
1/16 narrow
2. Duplicate genes or gene function are common for genes that encode essential cellular
functions.
3.
VI. Variations in expression
Penetrance: Looking at one gene …… the percent of individuals in a
population with a given genotype who exhibit the phenotype associated with the genotype.
Expressivity: Looking at one gene …… the extent to which a particular
genotype is expressed phenotypically in one individual.
Penetrance and expressivity can be affected by modifier genes, epistatic genes, and
environmental effects.
Lecture Syllabus Page | Home Page
This web site created and maintained by Dr. Laura
Runyen-Janecky
Last updated on 19 Jul 1999

